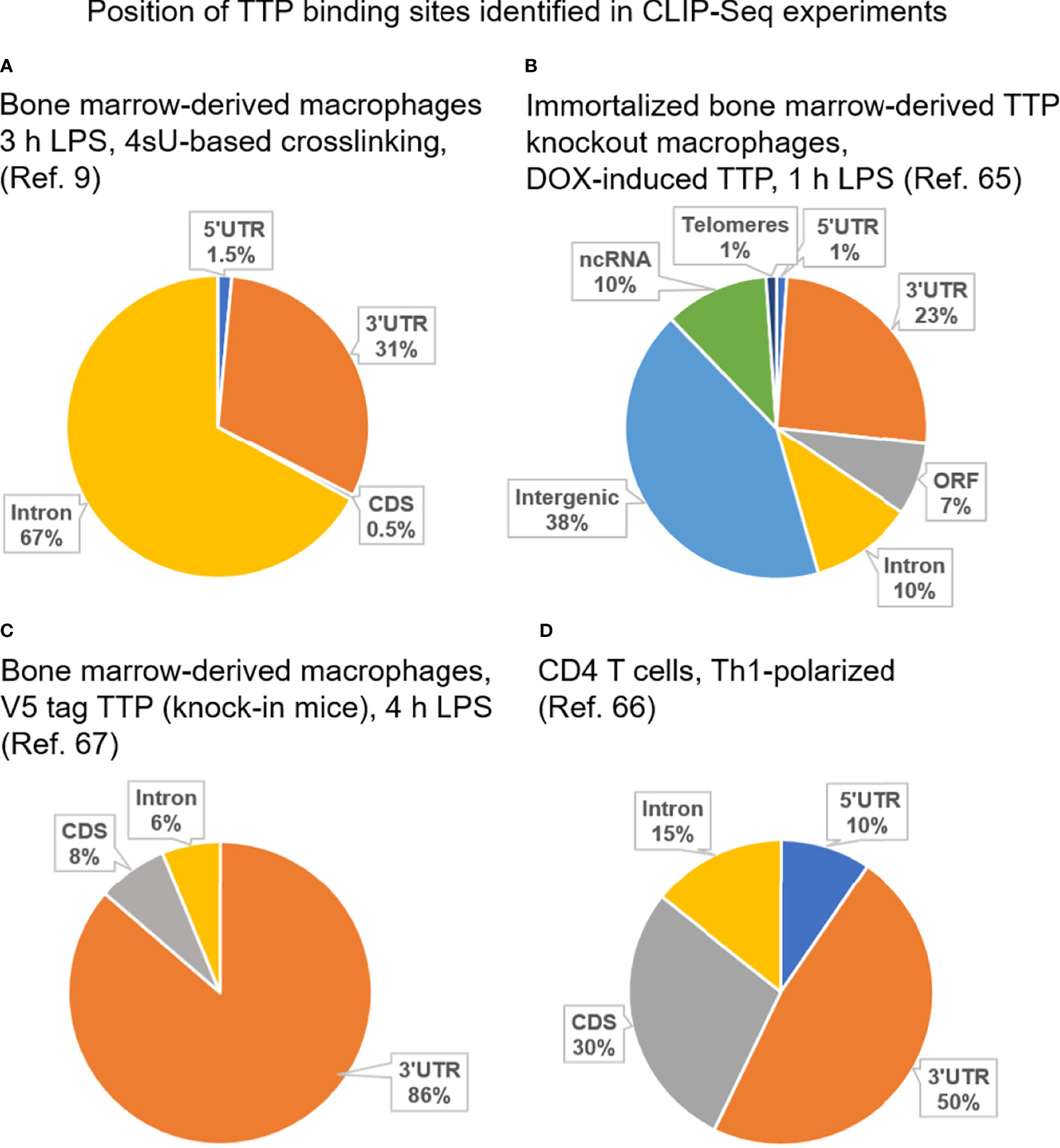
Frontiers | Conceptual Advances in Control of Inflammation by the RNA-Binding Protein Tristetraprolin

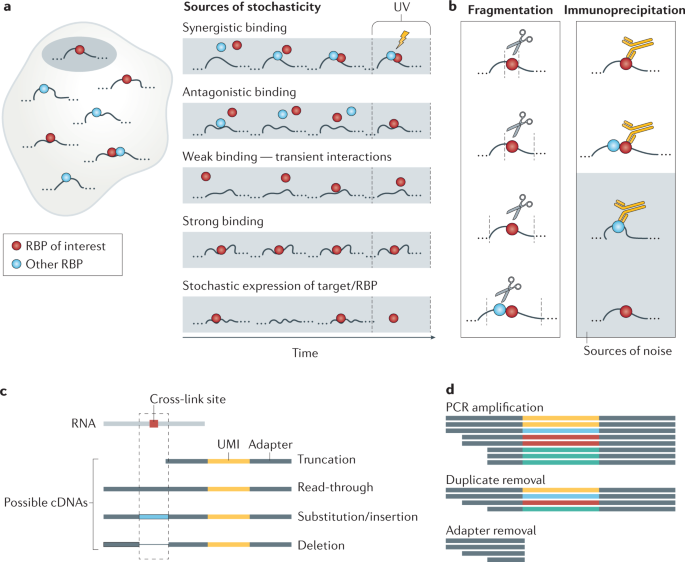
Advances in the characterization of RNA‐binding proteins - Marchese - 2016 - WIREs RNA - Wiley Online Library

Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell

Computational approaches for the analysis of RNA–protein interactions: A primer for biologists | Semantic Scholar

Uncovering the RNA-binding protein landscape in the pluripotency network of human embryonic stem cells - ScienceDirect

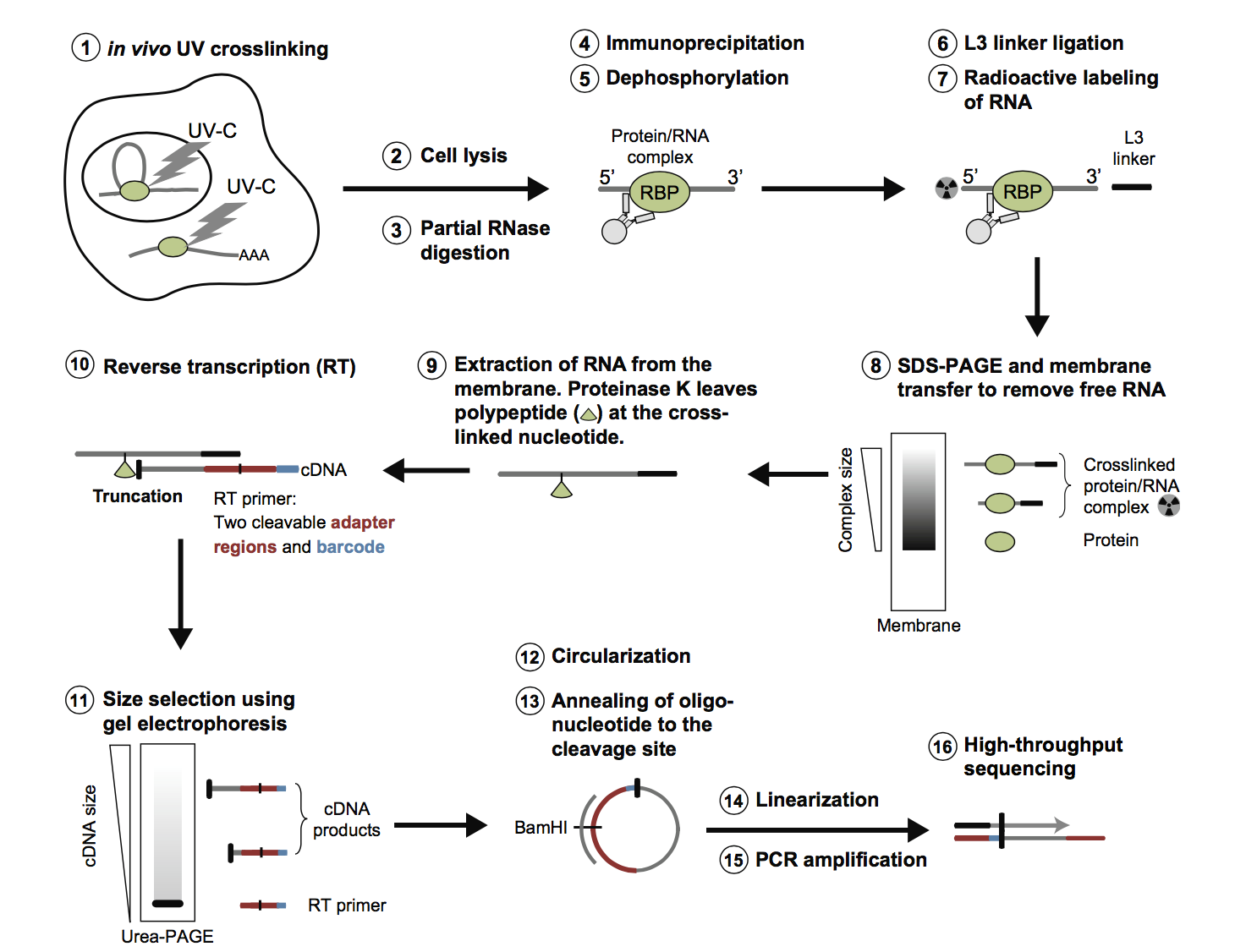
iCLIP - Transcriptome-wide Mapping of Protein-RNA Interactions with Individual Nucleotide Resolution | Protocol
Analysis of the nucleocytoplasmic shuttling RNA-binding protein HNRNPU using optimized HITS-CLIP method | PLOS ONE
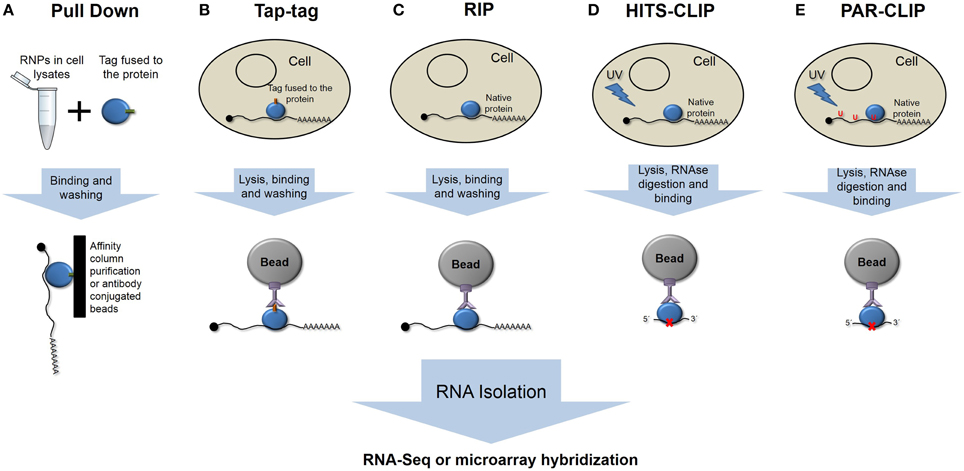
Frontiers | Stem Cell Ribonomics: RNA-Binding Proteins and Gene Networks in Stem Cell Differentiation

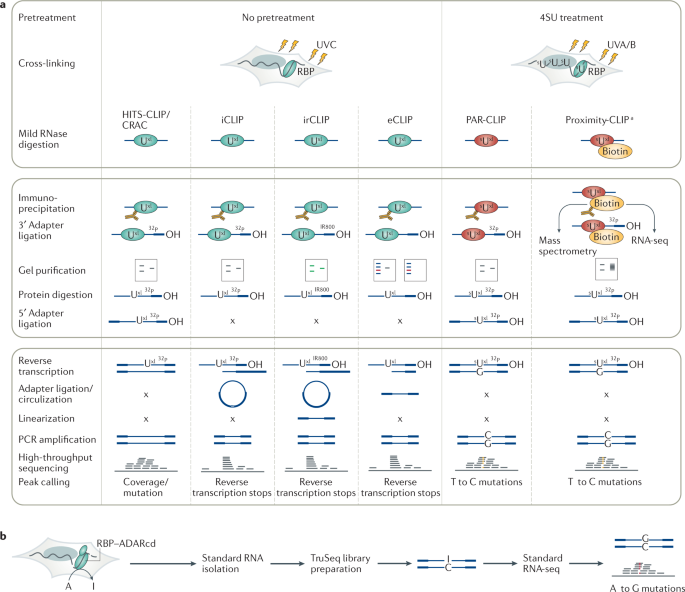
Outline of HITS-CLIP, PAR-CLIP and several variants, iCLIP, iCLAP and... | Download Scientific Diagram
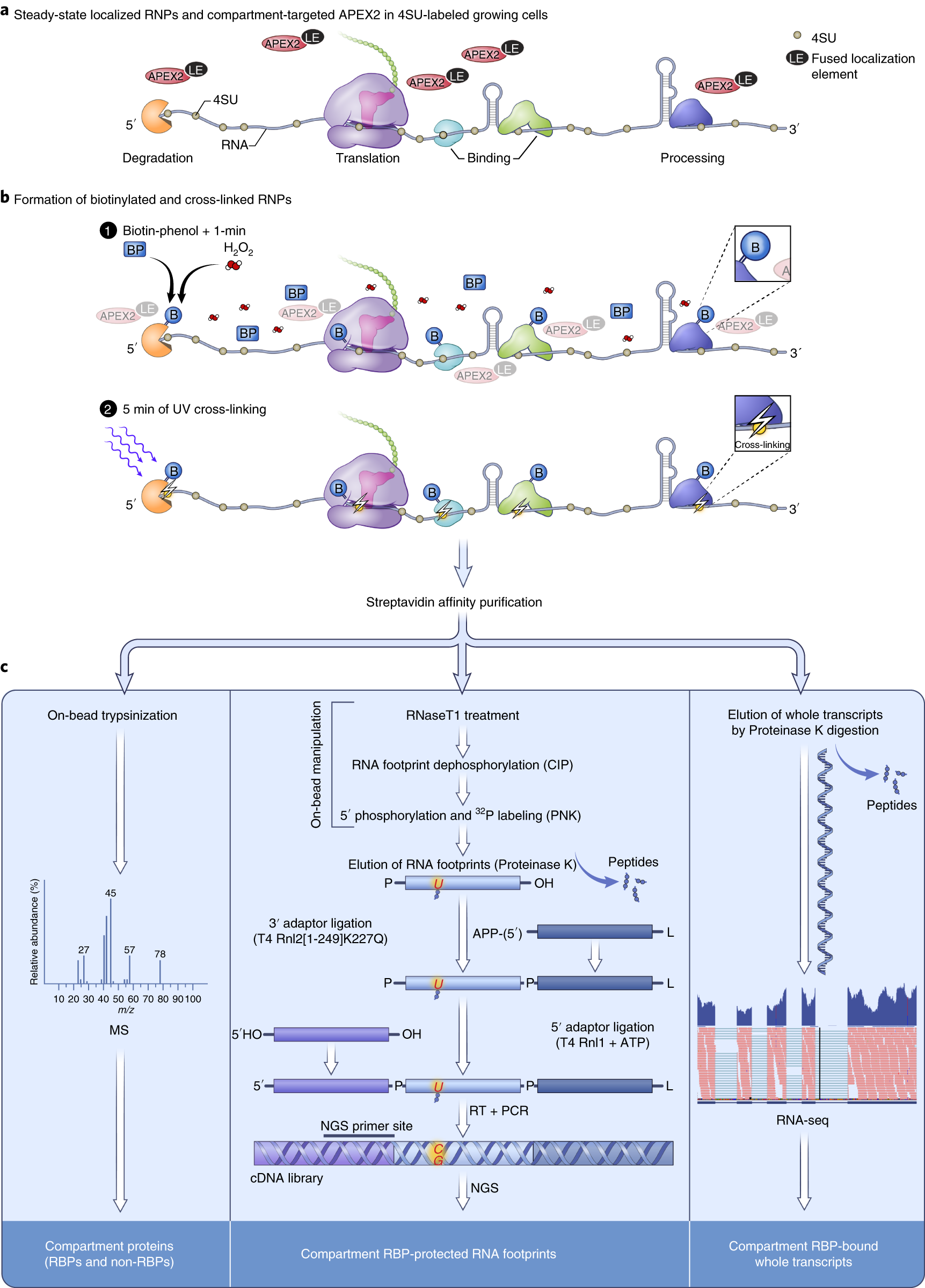
Proximity-CLIP provides a snapshot of protein-occupied RNA elements in subcellular compartments | Nature Methods

omniCLIP: probabilistic identification of protein-RNA interactions from CLIP-seq data | RNA-Seq Blog

RNA structure, binding, and coordination in Arabidopsis - Foley - 2017 - WIREs RNA - Wiley Online Library
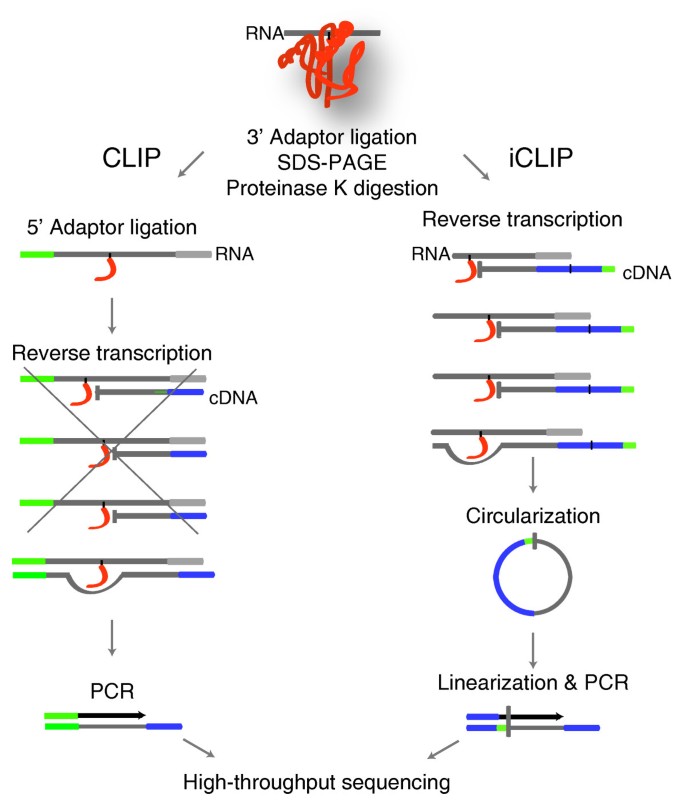
Analysis of CLIP and iCLIP methods for nucleotide-resolution studies of protein-RNA interactions | Genome Biology | Full Text
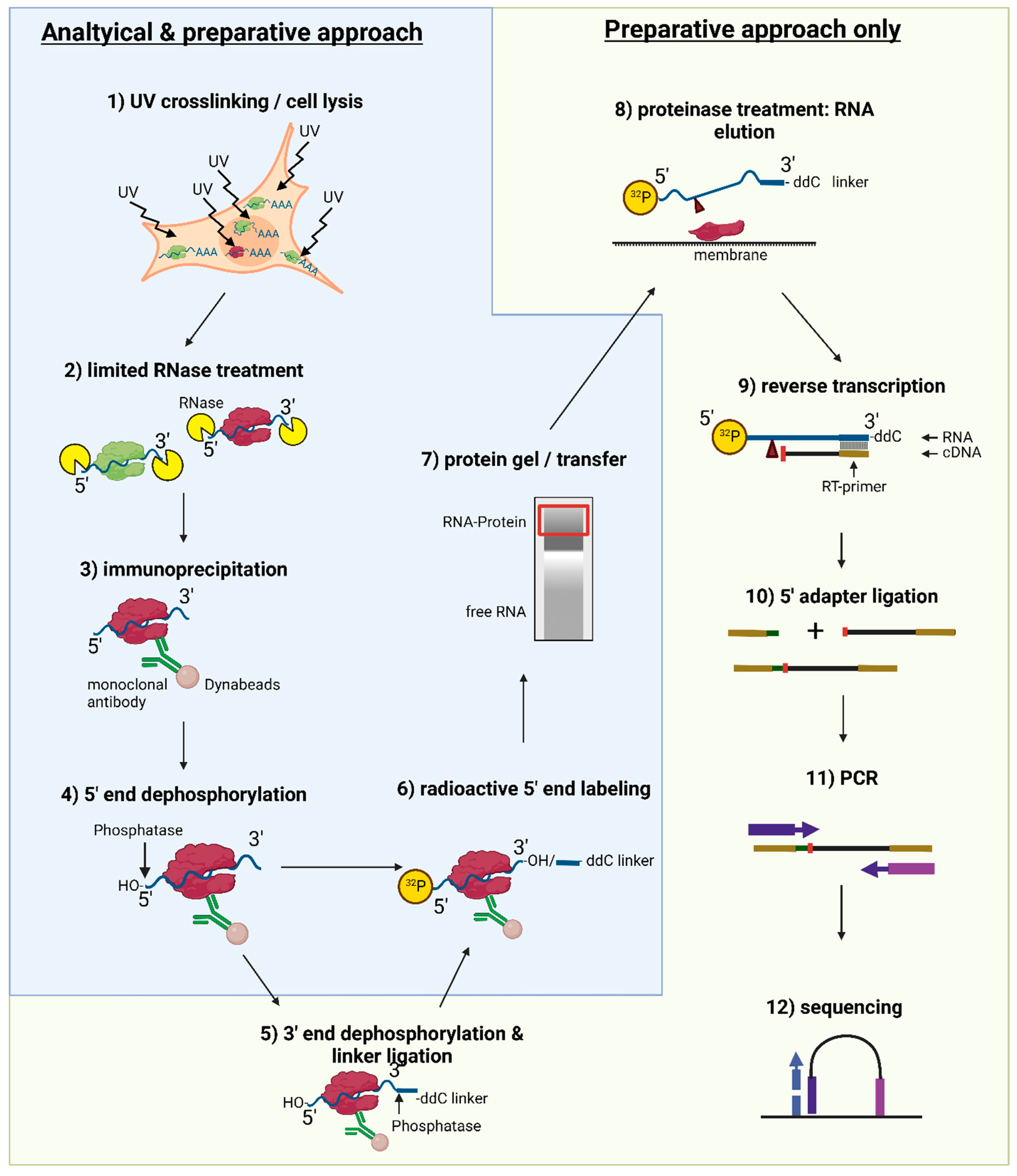
MPs | Free Full-Text | Studying miRNA–mRNA Interactions: An Optimized CLIP-Protocol for Endogenous Ago2-Protein


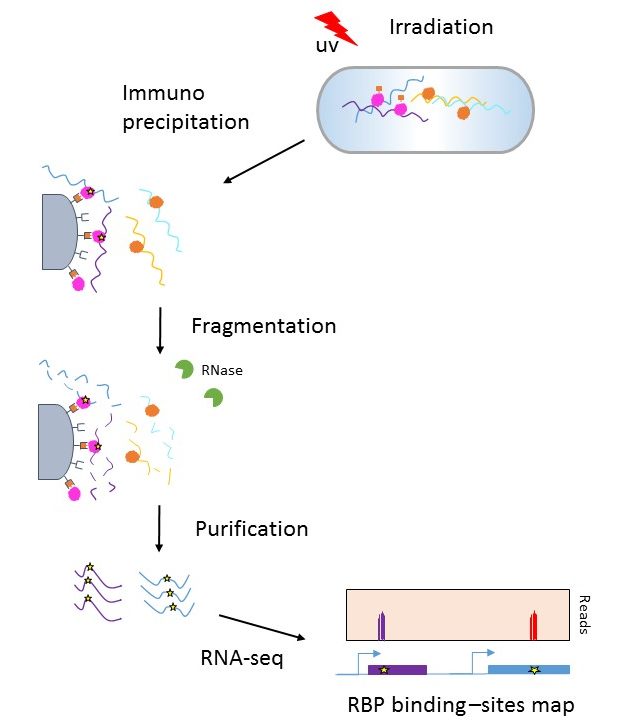

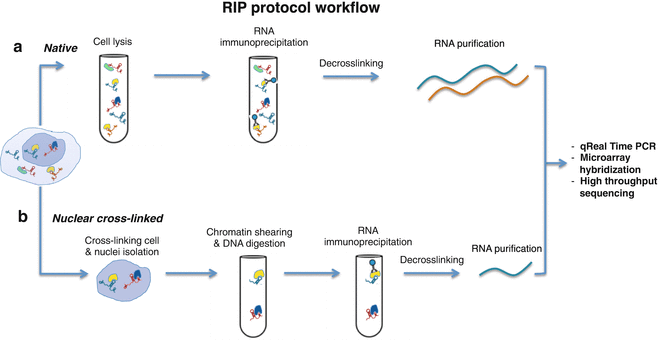


-2.jpg)